Here we post some of the cool microscopy images and videos we made.
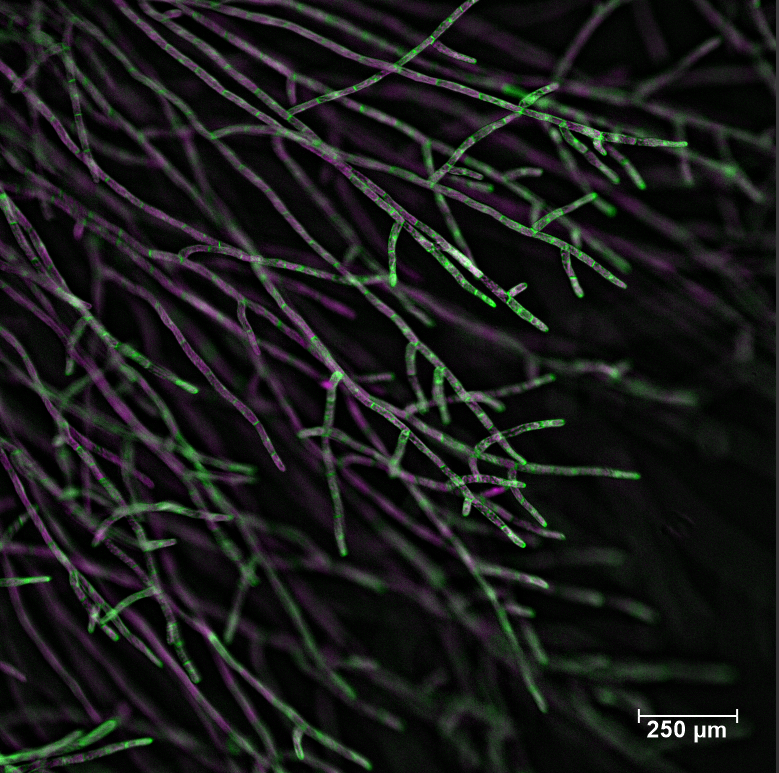
These are moss (Physcomitrium patens) protonema filaments stained with calcofluor white. Calcofluor white is a stain that labels the plant cell wall, and its fluorescence is shown here in cyan. The chlorophyll autofluorescence is in yellow/orange. Cells with fewer and smaller chloroplasts and oblique adjoining cell walls are caulonema cells, which grow faster and spread the plant. Cells with many large chloroplasts visible towards the bottom of the image are called chloronema cells. These cells are primary sites for photosynthesis. This is a deconvolved maximum Z projection image made on Olympus FV3000 confocal using the UPlanSAPO 10x/0.40 NA objective and high sensitivity detectors.
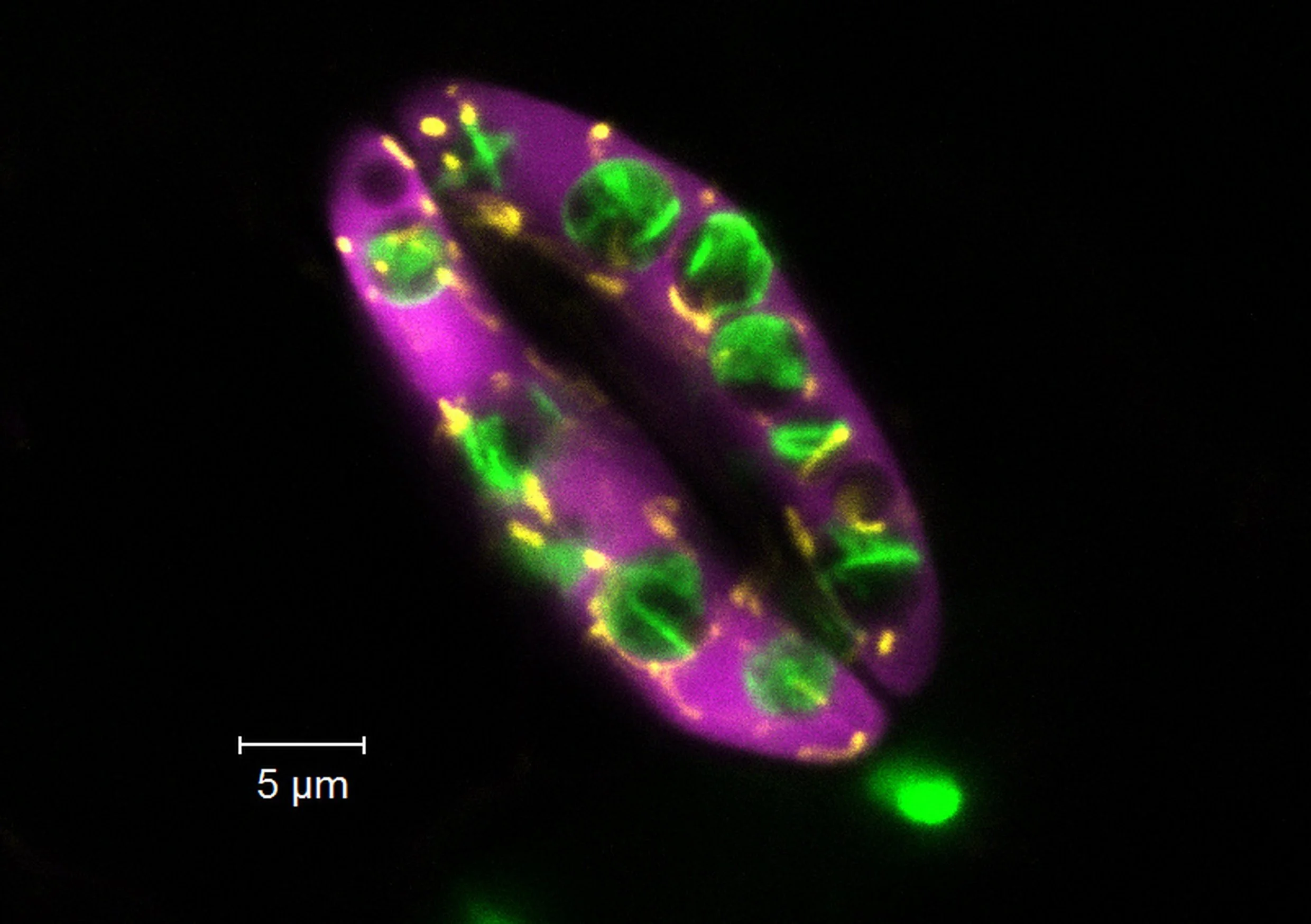
Moss (Physcomitrium patens) protoplasts (wallless cells) transformed with chloroplast envelope protein tagged with mGFP (green). The chlorophyll autofluorescence is in magenta. The protrusions of the chloroplast envelope are called stromules. This is a maximum-intensity projection along Z from a deconvolved stack captured on a confocal FV3000 (Evident/Olympus) microscope with a 60x water (NA 1.2) objective.
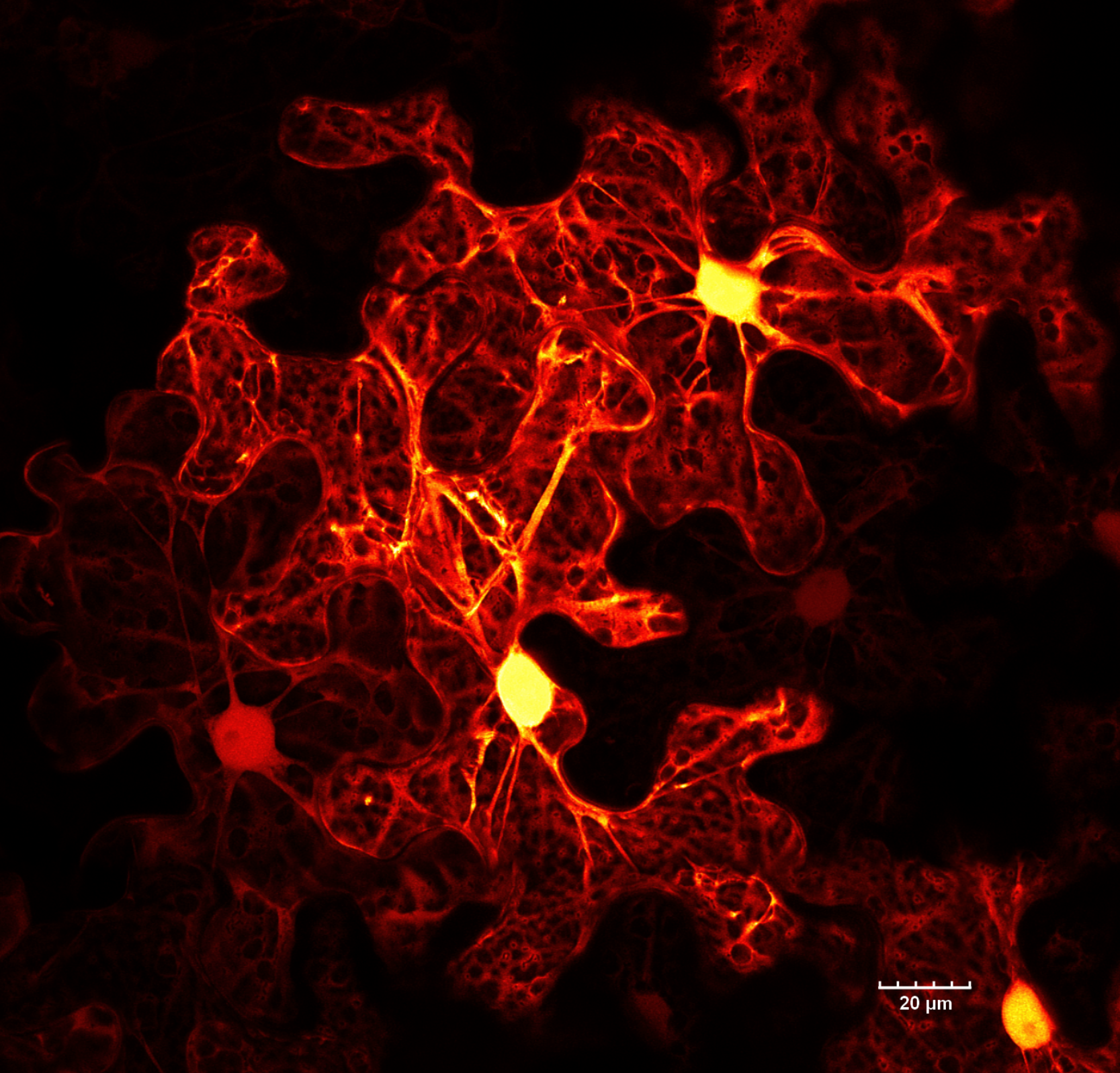
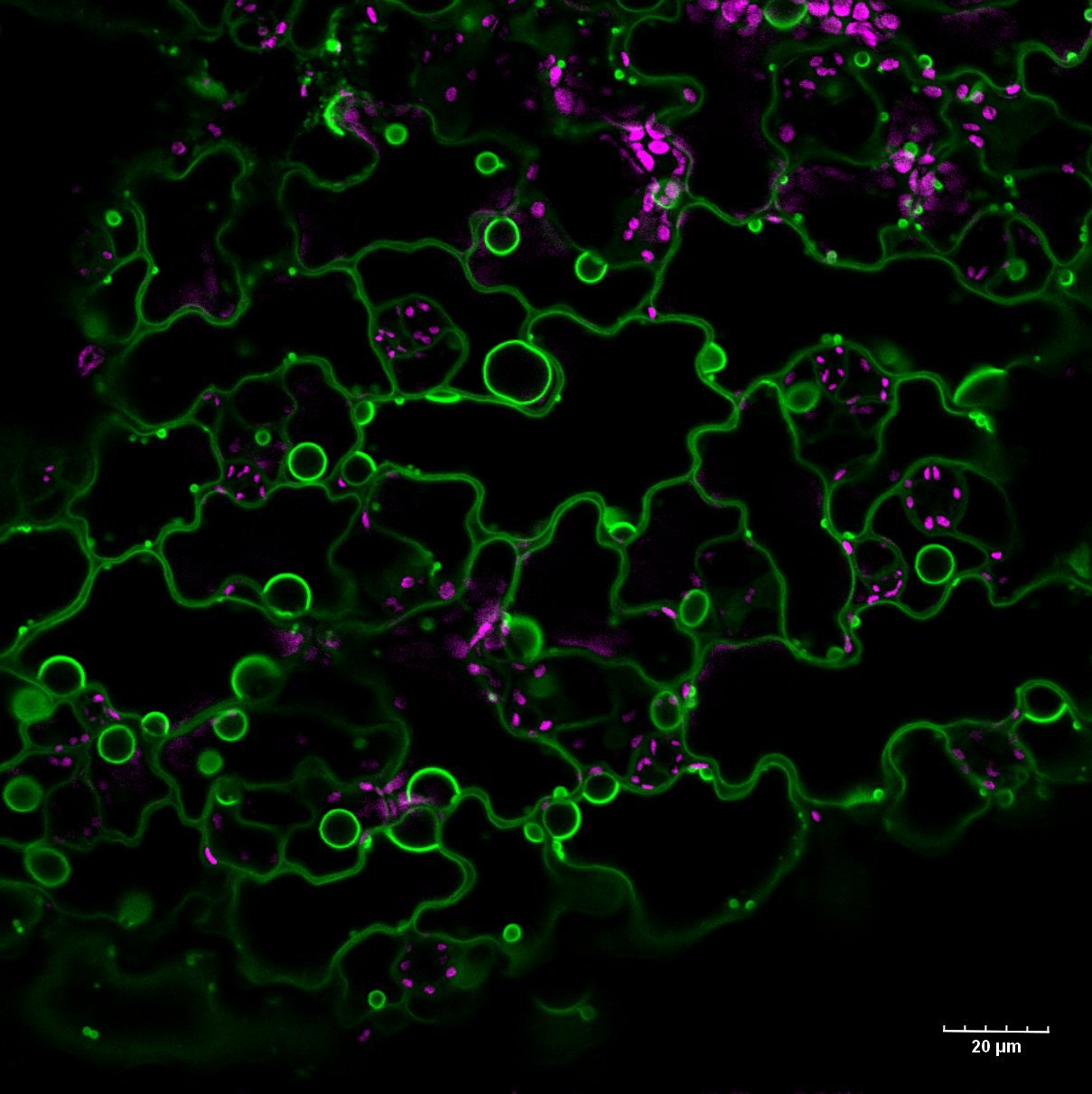
These epidermal cells of tobacco (Nicotiana benthamiana) leaf are expressing cytosolic GFP. The brightest region is the nucleus, which is surrounded by a large central vacuole. The cytosol is restricted to the cell periphery. Transvacuolar strands connect the nucleus with other parts of the cell.
This is a section of juvenile moss (Physcomitrium patens) tissue, called protonemata. The apical cells expand by tip-growth, while subapical cells sometimes divide to make lateral filaments. Cytosolic GFP is in green, and chlorophyll autofluorescence is in magenta.
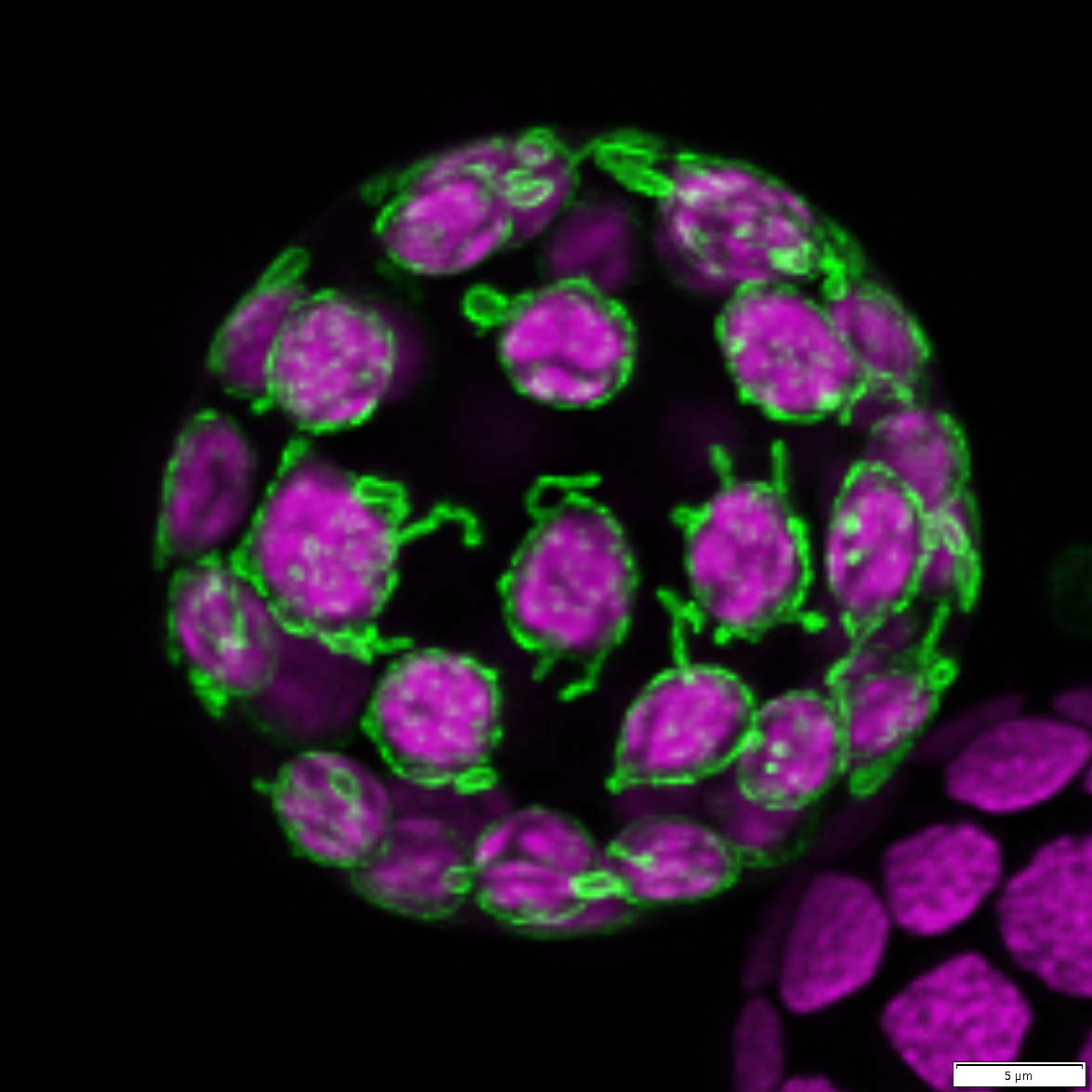
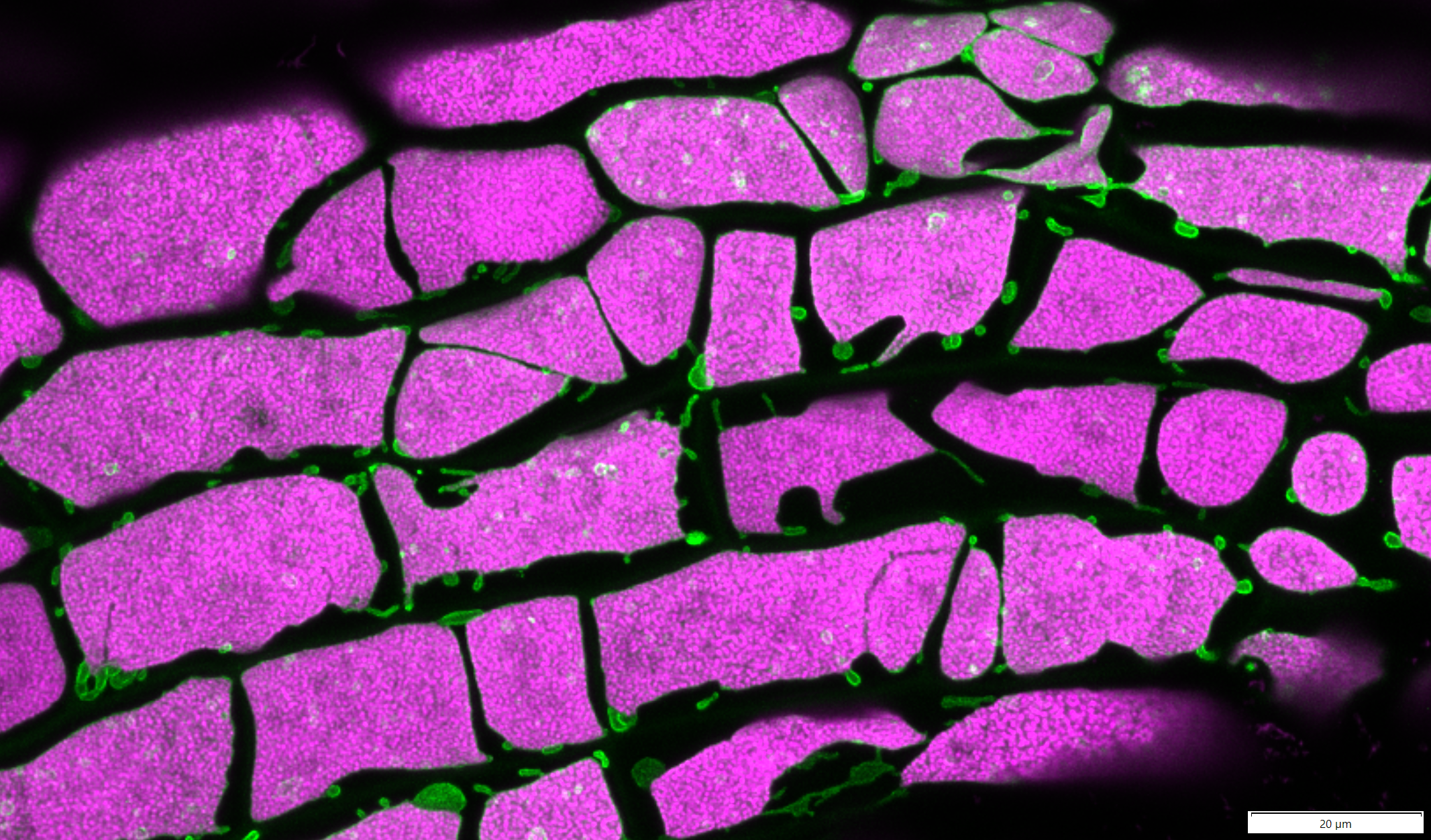
A top section of moss Physcomitrium patens gametophore comprised of “leaves” or phyllids. This gametophore was not stained or labeled. We are only seeing autofluorescence of numerous chloroplasts filling the cells. This is a maximum projection of 12 Z-stacks that have been subjected to deconvolution and stitched together. Z-stacks were captured on the Olympus FV3000, 10x/0.4 objective.
These are two Arabidopsis guard cells, with their cytosol in magenta, mitochondria in yellow, and chlorophyll autofluorescence in green. Guard cells form the stomata pore, a microscopic opening on the surface of leaves and stems. This is the main site of gas exchange (O2 and CO2) and transpiration (water release). The remaining epidermal cells surrounding the guard cells are not visible here.
A few cells from a “leaf” or phyllid from moss Physcomitrium patens gametophore. Cells are expressing a green fluorescence marker that is labeling the vacuolar membrane. Chloroplasts are also visible (in magenta) due to the autofluorescence of chlorophyll. This is a maximum projection of a Z-stack captured on the Olympus FV3000, 60x/1.2 W objective
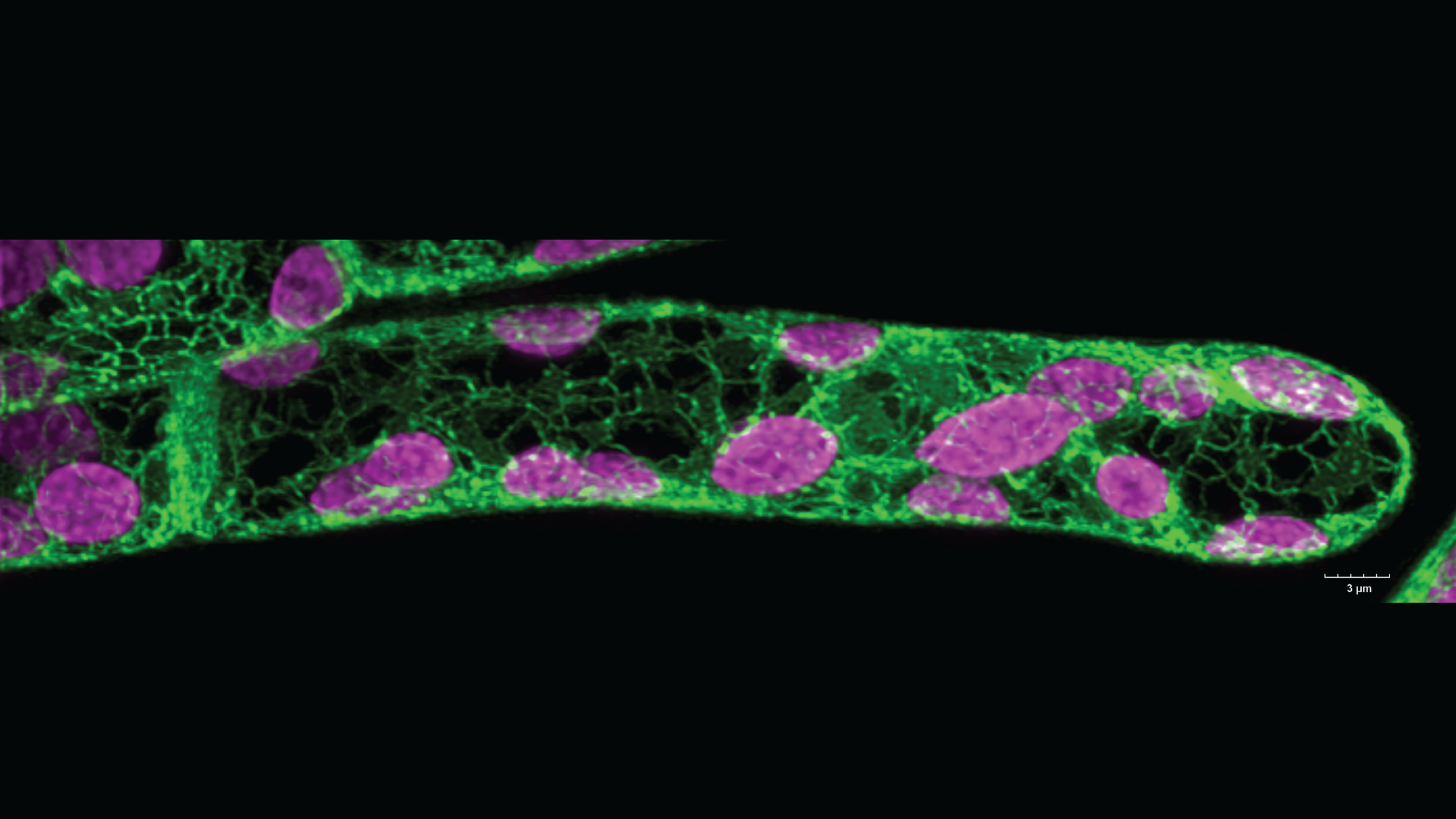
An undergraduate student in the lab, Andy Marchant, made this awesome image of a moss (Physcomitrium patens) apical chloronema cell. This is a beautiful example of how the ER network looks in plant cells (in green). The chlorophyll autofluorescence is in magenta.
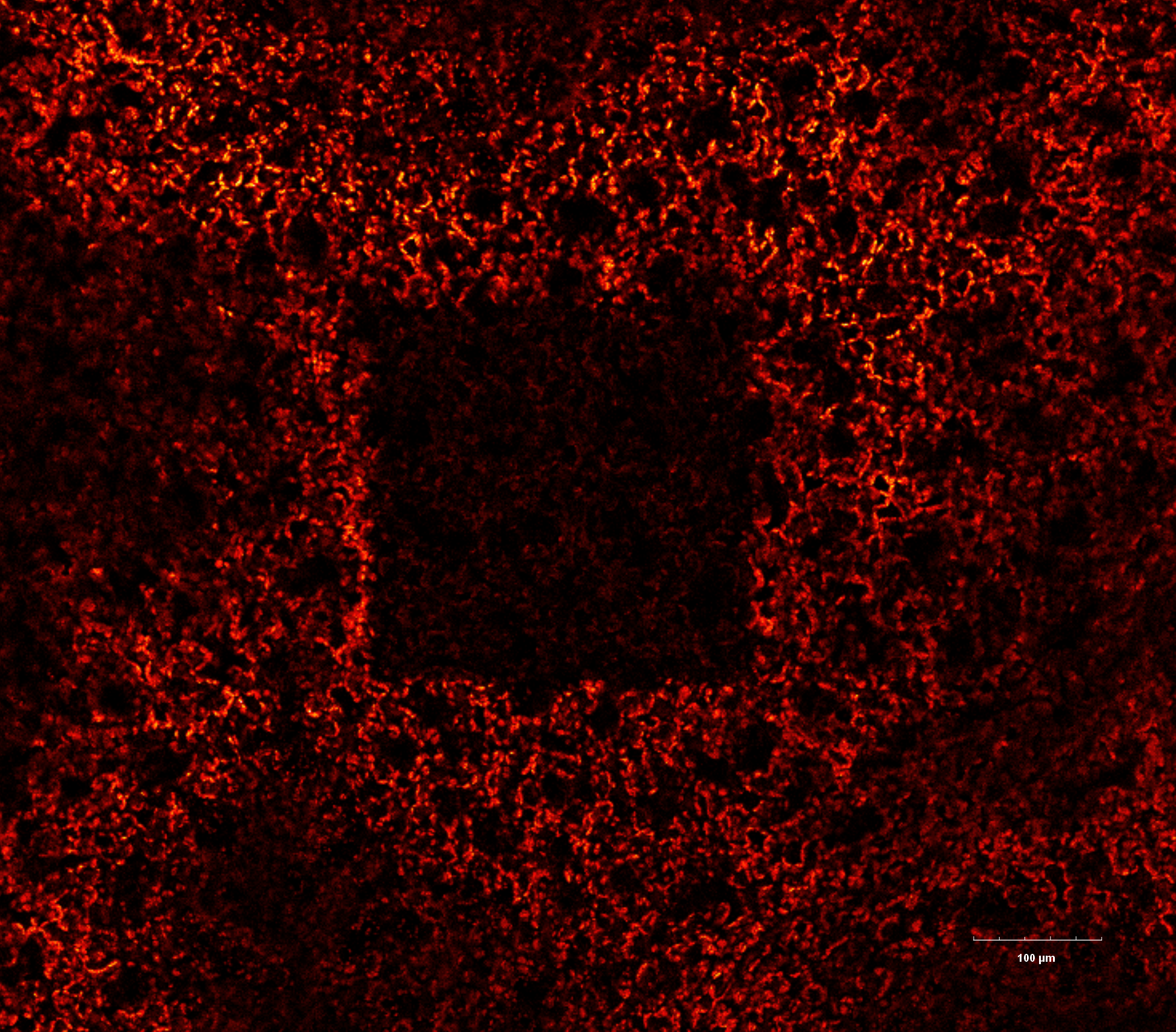
We were testing lasers and the spectral ability of our confocal (FV3000) on a zoomed-in section of a Brassica rapa leaf. But when we zoomed out, we saw that the lasers we used photobleached chlorophyll autofluorescence in the square area we had zoomed in on. We used a 10x/0.4 objective for this.
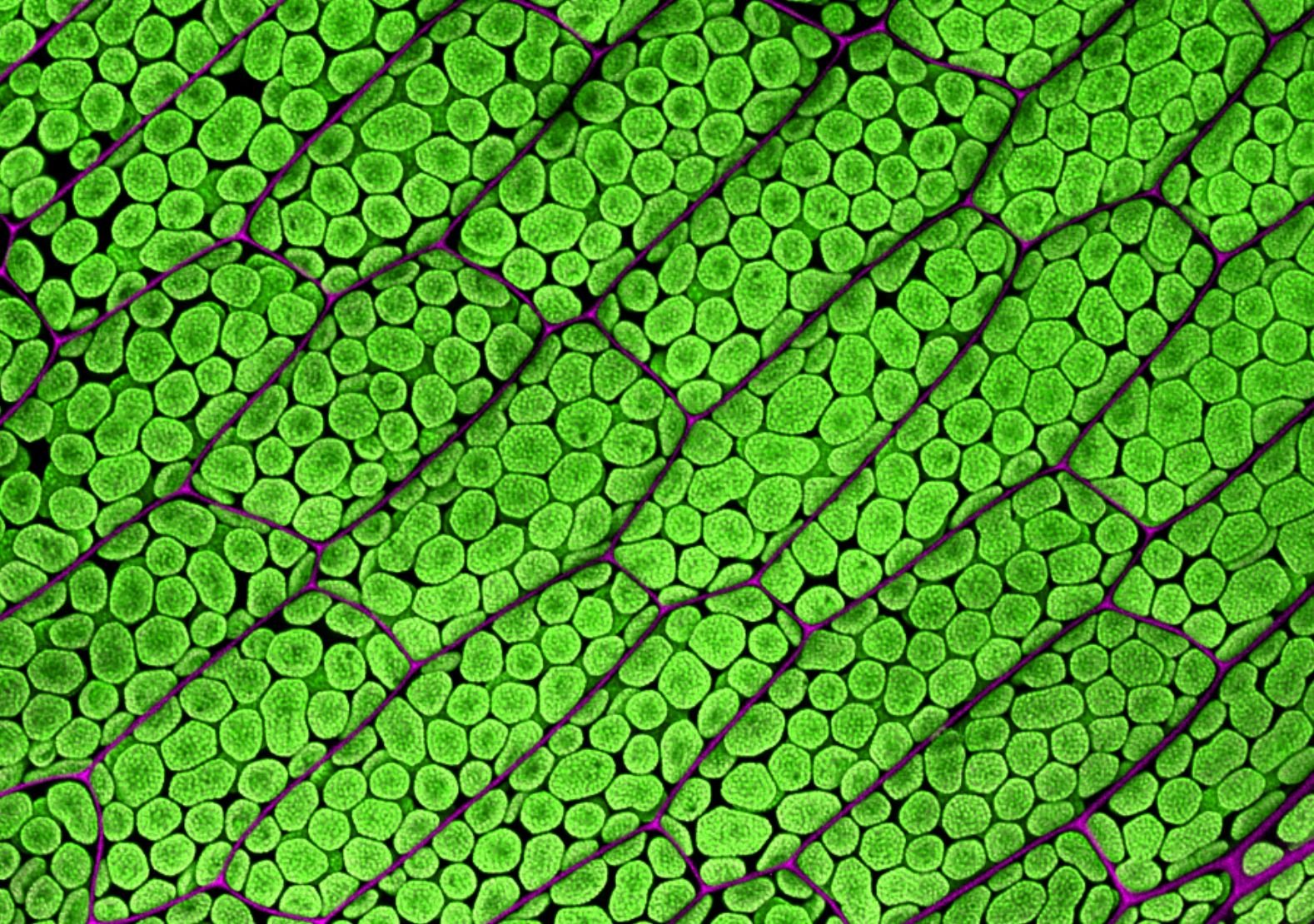
Surface (epidermal) cells of many plants are shaped like jigsaw puzzle pieces. 🧩 These are epidermal cells of tobacco (Nicotiana benthamiana) expressing cytosolic EGFP. The bright spherical structures are cell nuclei. While the epidermis is a continuous layer, the leaf's uneven surface and differences in cell expression levels give it the appearance of multiple layers. This stitched image (multiple fields of view merged together) was captured on an FV3000 confocal microscope from Evident/Olympus.
This is what you see when you magnify the edge of a moss (Physcomitrium patens) plant. The longer and relatively straight filaments are comprised of caulonema cells. These cells have fewer and smaller chloroplasts, oblique adjoining cell walls, and grow fast spreading the plant. At the bottom of the image, a few chloronema cells are visible. These cells are characterized by perpendicular adjoining cell walls and many large chloroplasts filling the cell volume. The cells in the image are stained with calcofluor white, which stains the cell walls; here shown in magenta. The chlorophyll autofluorescence is shown in green. This is a maximum projection of a deconvolved Z-stack captured on FV3000 confocal from Olympus, using the UPlanSAPO 10x/0.40 NA objective and high-sensitivity detectors.
Two apical caulonemal cells from moss, Physcomitrium patens stained with MDY64. MDY64 stains plasma and vacuolar membranes as well as other cytosolic compartments. Vacuoles are easily distinguished as empty black spaces surrounded by a single membrane. The only exception is the circular space in the middle of the cell, where the nucleus resides. MDY64 fluorescence is shown in Fall LUT. We can also see chloroplasts (in cyan) due to the natural autofluorescence of chlorophyll. Captured on Olympus FV3000 confocal with 60x/1.2 W objective and subjected to deconvolution.
Here is a fluorescent image of moss gametophore "leaf" cells. In the image, we can see chloroplasts (in green) and walls between neighboring cells (in magenta). Both are visible due to their natural autofluorescence. This is a deconvolved maximum Z projection image made on Olympus FV3000 confocal microscope from Olympus.
Haunted Chlamy A Z-stack through a part of the chloroplast from a single Chlamydomonas reinhardtii cell. The signal comes from chlorophyll autofluorescence. The image was captured on AXR confocal from Nikon using 100x/1.4 oil objective.
This is a single caulonemal filament from moss Physcomitrium patens stained with Calcofluor white. Calcofluor white stains the cell wall and here it’s shown in cyan. Only magenta chloroplasts are visible in the cell interior due to chlorophyll autofluorescence. Only the apical cell grows by adding new material to the tip, however, the cells further down the filament sometimes divide and create new growing cells and branching filaments. This is a maximum projection of a Z-stack captured on the Olympus FV3000 using a 20x/0.75 objective
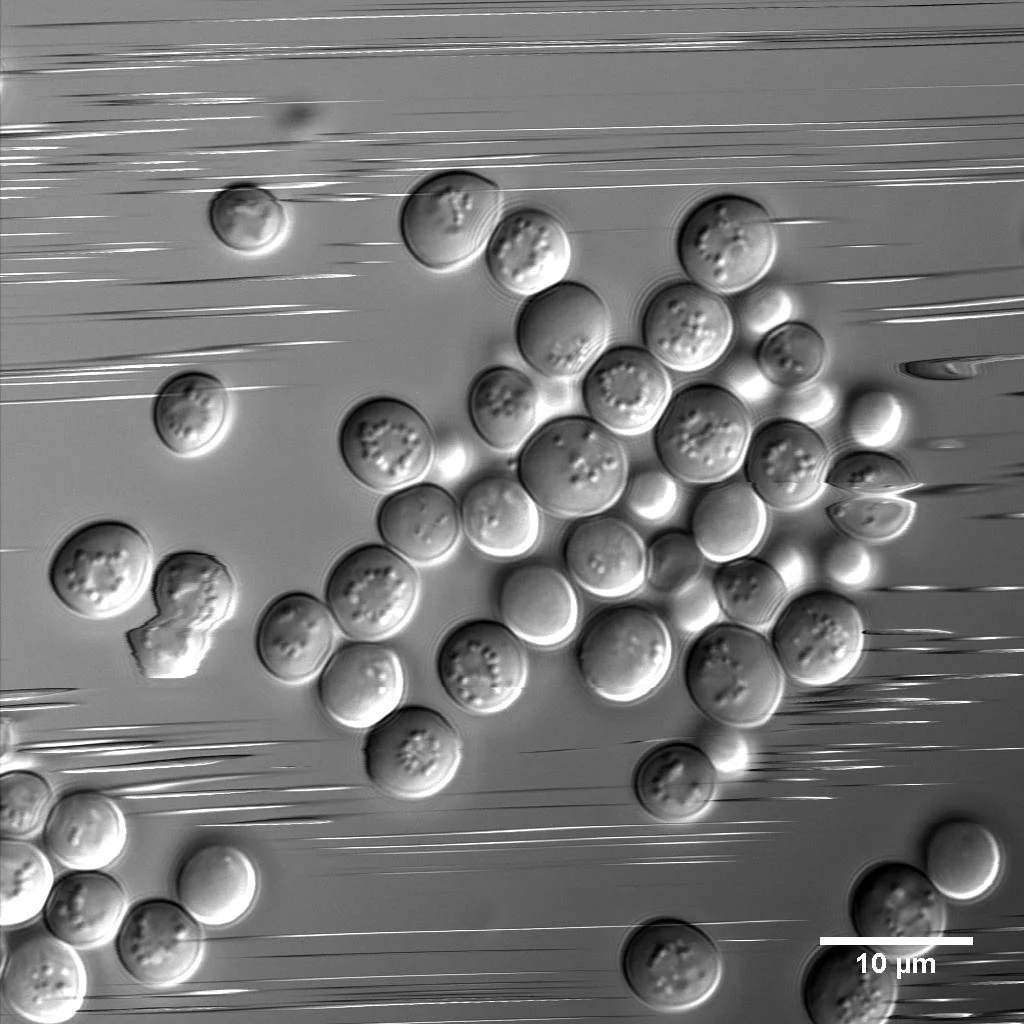
These Brewer's yeast cells were in a hurry! This was captured with a point-scanning confocal microscope, which scans one point (pixel) at a time, moving from left to right and top to bottom. As a consequence, cells moving faster than the scan speed appear as streaks or exhibit distortions.
This is a section of the cotyledon (first leaf) surface from the flowering plant Arabidopsis thaliana, which is expressing the green fluorescent protein vacuolar membrane marker (vac-GB). Chloroplasts are in magenta, visible due to the autofluorescence of the chlorophyll. Captured on Olympus FV1000 confocal.
Did these chloroplasts have a seance! 👻🎃 These five Arabidopsis chloroplasts were extending their stromules (protrusions of the chloroplast envelope and stroma) towards each other. Stromules are visible due to the stroma-targeted fluorescent marker (in green). Stromules are comprised of both envelope membrane and stroma, but not chlorophyll-containing thylakoid membranes, visible here in magenta due to autofluorescence. The image was captured on an FV1000 confocal from Olympus.
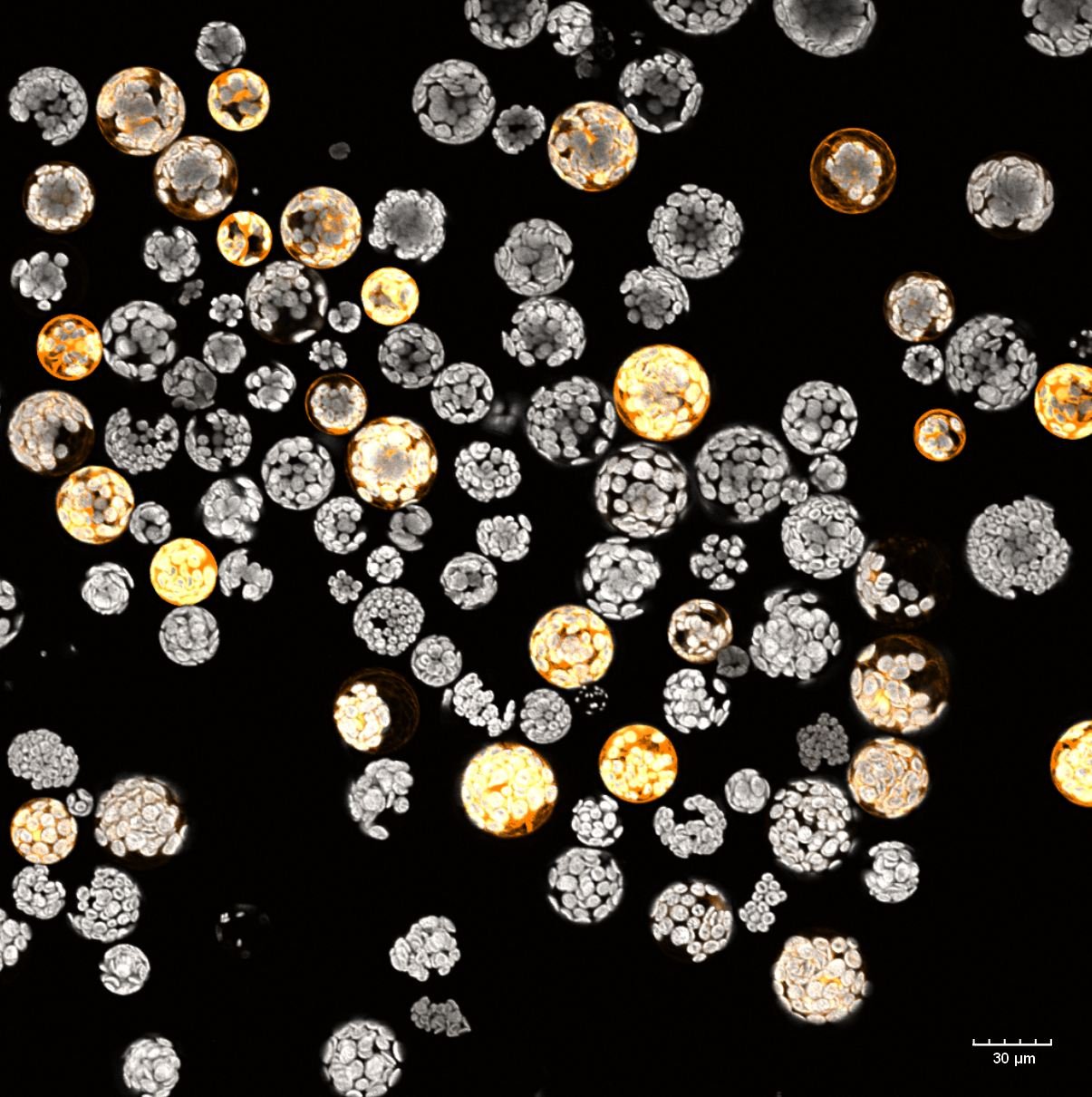
Protoplasts, plant cells without cell walls, isolated from the flowering plant Arabidopsis thaliana. The protoplast interior is filled with chloroplast visible due to their natural autofluorescence (in grey). Some protoplasts are expressing cytosolic GFP (green fluorescence protein). GFP fluorescence is shown in orange. This is a maximum projection of a Z-stack captured on the Olympus FV3000, 20x/0.75 objective.
This is what happens to the moss (Physcomitrium patens) chloroplasts when we grow cells on β-lactam antibiotics. β-lactams inhibit the synthesis of the peptidoglycan in the chloroplast envelope, leading to their dramatic expansion. Magenta - chlorophyll autofluorescence, Green -mGFP-tagged protein targeted to the chloroplast envelope. The envelope can produce tubular protrusions called stromules, many of which are visible here. These are moss gametophore cells, many of which have only one giant chloroplast. This is a deconvolved Z-maximum projection of an image captured on the FV3000 confocal microscope from Olympus using a 60x/1.2W objective.
These are filaments of the Charophyte algae Klebsormidium nitens. We only see chloroplasts due to chlorophyll autofluorescence. Each cell has one elliptical or discoid-shaped chloroplast. This is a maximum projection of a Z-stack, captured on Olympus FV3000 confocal with 60X/1.2W objective.
This is an edge of a mature Arabidopsis root cell. The vacuolar membrane (green) is pressed against the plasma membrane (magenta). Cytosol and most other organelles have to fit in the tight space in between. Both membranes were labeled with fluorescent proteins. The image was subjected to deconvolution. The image was captured on Olympus FV3000 confocal with a 60x/1.2W objective.
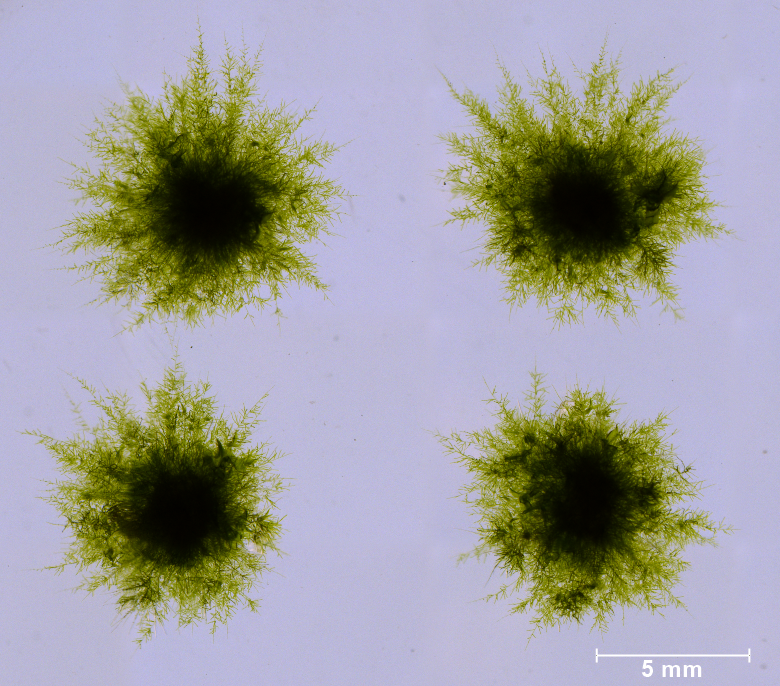
Four tufts of moss Physcomitrium patens juvenile protonemata.
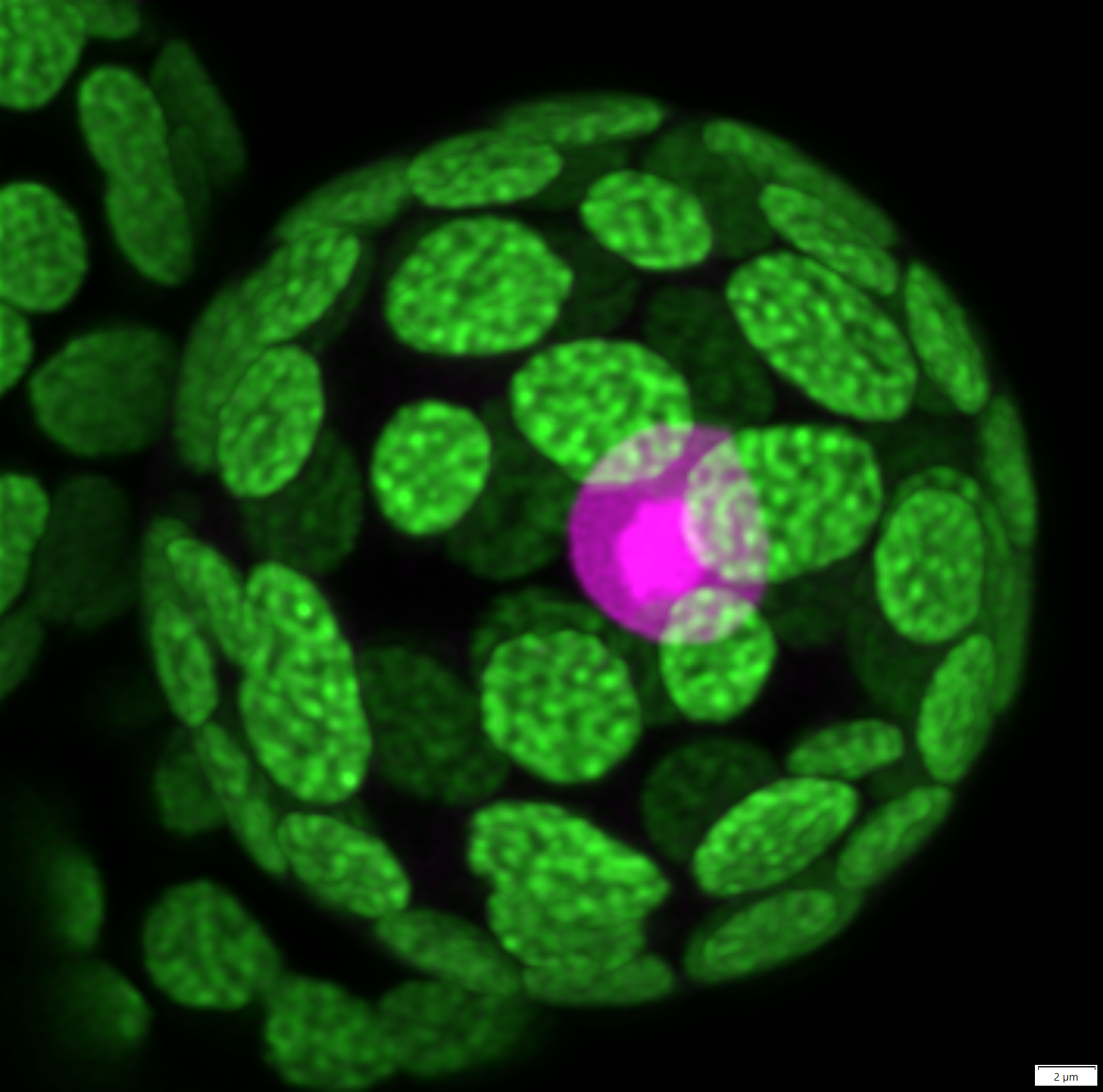
This is a moss (Physcomitrium patens) protoplast (wall-less cell) transformed with a nuclear marker (in magenta). The much brighter area is the nucleolus. The chlorophyll autofluorescence is in green. This is a deconvolved Z-maximum projection of an image captured on the FV3000 confocal microscope (Olympus) using a 60x/1.2W objective.
Different types of plastids in the leaf of the flowering plant Arabidopsis thaliana. Plastids are labeled with the stroma-targeted fluorescence marker RecARed. RecARed fluorescence is false-colored yellow/orange, while the chlorophyll autofluorescence is in grey. The plastids in the epidermal cells are smaller and accumulate much more of the RecARed marker hence are very bright. These plastids also produce a lot of string-like protrusion of their envelope, called stromules. The plastids from the deeper cells are bigger and have more chlorophyll compared to the RecARed marker so appear greyer. This is a maximum projection of a Z-stack captured on the Olympus FV3000, 60x/1.3 silicone objective.
These are filaments of the Charophyte algae Klebsormidium nitens. We only see chloroplasts due to chlorophyll autofluorescence. Each cell has one elliptical or discoid-shaped chloroplast. This is a maximum projection of a Z-stack, captured on Olympus FV3000 confocal with 60X/1.2W objective.
The image shows how chloroplasts (in magenta) indent the vacuole to make room for themselves. The vacuolar membrane is labeled with a GFP marker (green). These are moss (Physcomitrium patens) apical and subapical chloronema cells. This is a deconvolved maximum projection of a Z-stack, captured on Olympus FV3000 confocal with 60X/1.2W objective.
Tobacco (Nicotiana benthamiana) leaf was injected with the bacteria Agrobacterium tumefaciens. The bacteria transferred a piece of DNA with the GFP (green fluorescent protein) gene into plant cells, forcing them to make a lot of free protein. GFP fluorescence was excited by blue light.
This is a border of two Arabidopsis hypocotyl cells. The cell wall (invisible due to lack of fluorescence) is between two plasma membranes (magenta). The vacuolar membranes (green) press the cytosol and other organelles (including the chloroplast, in blue) against the PM. The two membranes are labeled with fluorescent proteins, while the chloroplast is visible due to the chlorophyll autofluorescence. The image was captured on Olympus FV3000 confocal with a 60x/1.2W objective.
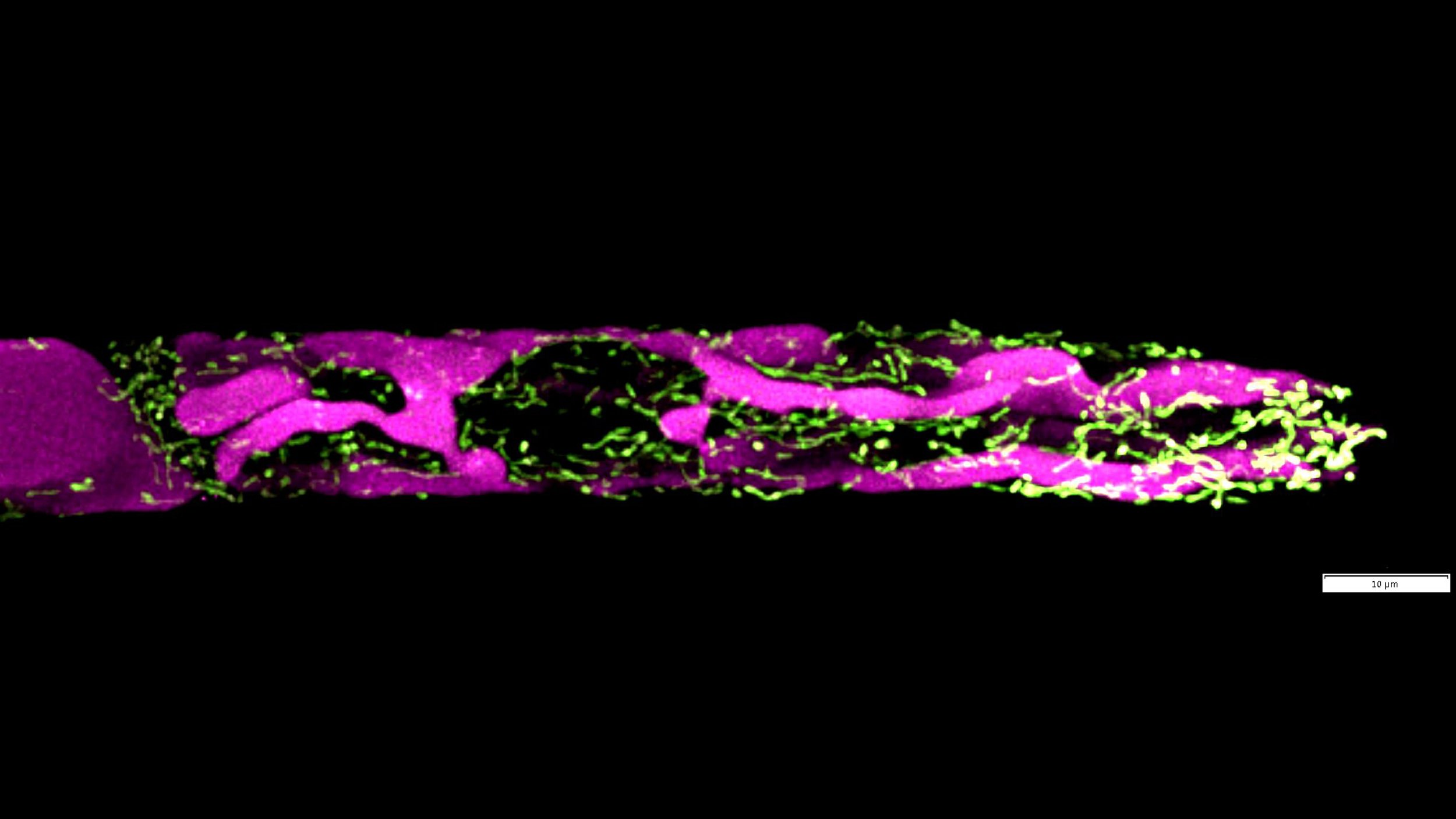
The image shows an apical caulonemal cell from moss Physcomitrium patens stained with BCECF, which is a ratiometric-based pH indicator. Due to their pH difference, vacuoles (low luminal pH) are more visible in the first channel (magenta), while what I assume are mitochondria are more visible in the second channel (green). Vacuoles form large tubules that run parallel to the longitudinal axis of the cell. Mitochondria form much smaller and shorter tubules distributed all over the cell. The image is a maximum projection of a deconvolved Z-stack, made on our Olympus FV3000 confocal from Olympus, using a Galvano scanner, high sensitivity detectors, and UPLSAPO60X water NA 1.2 objective.
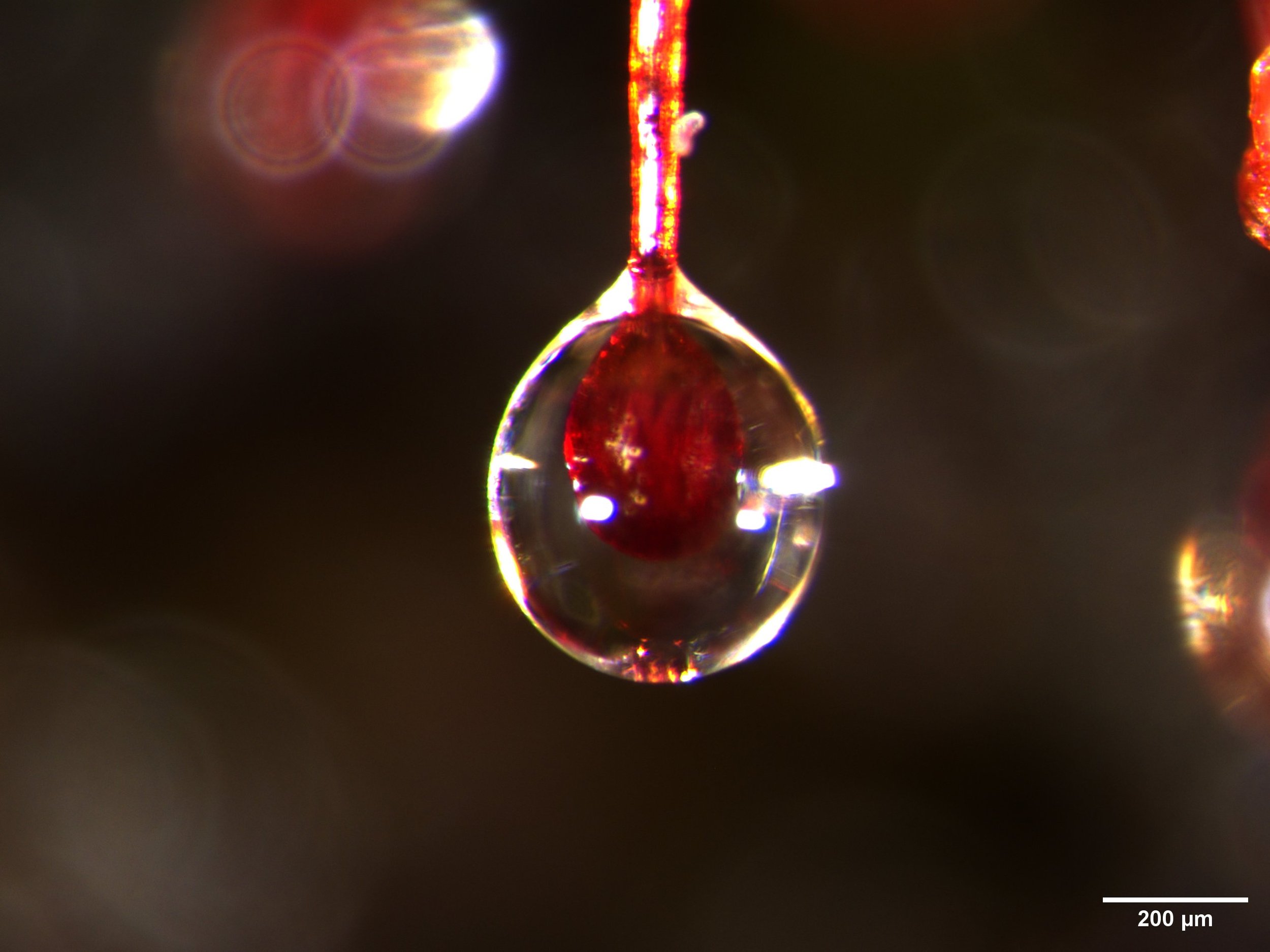
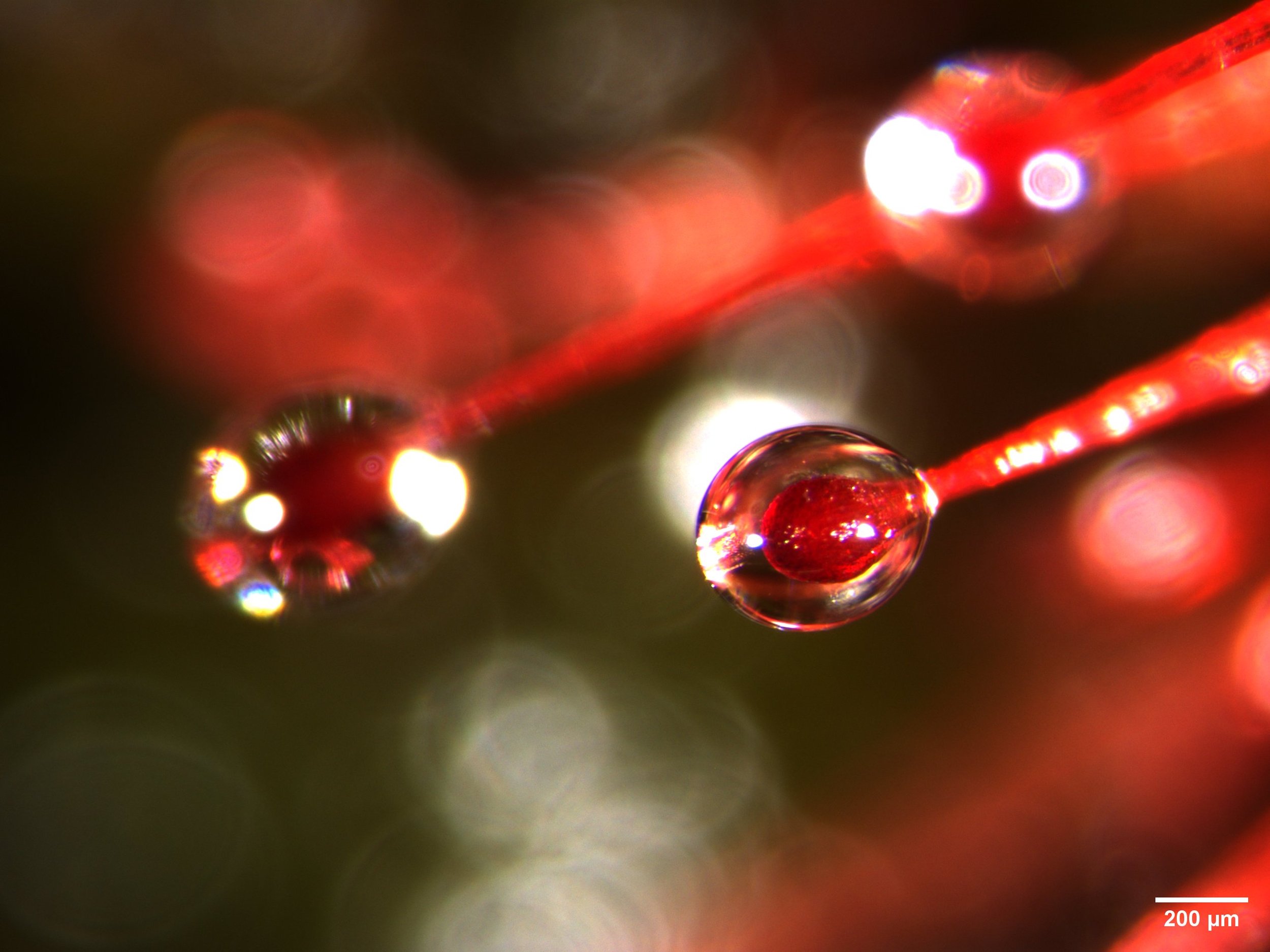
This is a close-up of mucilage-covered tentacle tips from the leaf of a carnivorous sundew plant (Drosera spatulata). See the previous post. The images were made with the THUNDER Imager Model Organism dissecting scope from Leica.
This is a close-up of mucilage-covered tentacle tips from the leaf of a carnivorous sundew plant (Drosera spatulata). See the previous post. The images were made with the THUNDER Imager Model Organism dissecting scope from Leica.
This is a carnivorous Sundew plant (Drosera spatulata). These amazing plants grow as small rosettes and their leaves are covered with tentacles that secrete mucilage on the tips. The plants use the mucilage to adhere the insect prey long enough for the tentacles and the leaf to curve and entrap the prey. These plants grow on nutrient-poor soil, so they have to supplement their diet with some insects.
This is what you see when you zoom in on fungus, growing uninvitedly on my "sterile" moss plates. What a beautiful hyphal network. The image was made with the THUNDER Imager Model Organism dissecting scope (M205FCA) from Leica, available at the WUSTL Biology department imaging core. The image is a stitched composite from 9 (3 by 3) tile images captured using the 1x/0.6 objective with a 6.3x mag and greyscale camera.
Different types of plastids in the leaf of the flowering plant Arabidopsis thaliana. Plastids are labeled with the stroma-targeted fluorescence marker RecARed. RecARed fluorescence is false-colored green, while the chlorophyll autofluorescence is in magenta. The plastids in the epidermal cells are smaller and accumulate much more of the RecARed marker hence appear green. These plastids also produce a lot of string-like protrusion of their envelope, called stromules. The plastids from the deeper cells are bigger and have more chlorophyll compared to the RecARed marker. This is a maximum projection of a Z-stack captured on the Nikon AXR confocal, 40x/1.15 water objective.
This is a single gametophore with the accompanying rhizoids from moss Physcomitrium patens. The image shows a single green gametophore from moss Physcomitrium patens. Gametophore is composed of many interlaced leaf-like structures, phyllids. Long and brownish rhizoids are protruding from the base of the gametophore. The image was made with the Olympus SZX7 dissecting scope with a DP71 camera from Olympus.
A collage of individual 7-day-old moss (Physcomitrium patens) plants, regenerated from a single protoplast (plant cell with removed cell walls). When moss cells are protoplasted, they behave like spores and can rebuild the whole plant. Over four days, protoplasts, when cultured on osmotically supported media, regenerate their cell walls and differentiate into chloronema cells, which slowly start to grow and divide. When moved to regular media, the plants start to rapidly grow and divide. Within 3 days plants expand from 1-2 cells to what you see in the image. At one point, some of the chloronema cells start to differentiate into caulonema cells, both of which are characterized by tip growth The images were made with the THUNDER Imager Model Organism dissecting scope (M205FCA) from Leica using the 1x/0.03 objective with a 4x mag, greyscale camera, and below the sample illumination. The collage was made in Illustrator.
Not all microscopy images turn out great. Nevertheless, they can still be cool. This one definitely has an abstract art vibe. The image shows Zmax projection of tobacco leaf, transiently overexpressing vacuolar membrane localized AtPIEZO1 tagged with EGFP. The EGFP fluorescence is in green. The leaf was also stained, unsuccessfully, with Mitotracker, in magenta. The chlorophyll autofluorescence is blue.
Our microscopy videos:
What happens when you poke a leaf of the carnivorous sundew plant?
A rapid calcium wave spread from the poked leaf to the whole plant. The plant is expressing the cytosolic calcium biosensor (GCaMP3), which allows us to see the changes in the cytosolic calcium levels. The change in color correlates with the change in calcium concentration. Burgundy - Low, Orange - Medium; Yellow - High Ca2+.
This project was a great collaboration with Carl Procko from the Chory lab at SALK. The video was captured with THUNDER Imager Model Organism dissecting scope from Leica, a part of WashU Biology imaging core.
Two videos show what happens when a fly lands on the leaf of the carnivorous sundew plant (left) or when you bend tentacles (right). The sundews leaf tentacles are mechanosensitive! They show rapid calcium transients in response to bending with a glass probe or stimulation by an insect. The calcium transient subsequently spread from the tentacle into the leaf surface.
The cells are expressing the cytosolic calcium biosensor (GCaMP3), which allows us to see the changes in the cytosolic calcium levels. The change in color correlates with the change in calcium concentration. Blue - Low, Green - Medium; Yellow - High Ca2+. The bright signal coming from the fly (Drosophila melanogaster) is autofluorescence.
The video shows the 3D reconstruction of the Z-stack image from different perspectives. Different types of plastids in the leaf of the flowering plant Arabidopsis thaliana were imaged. Epidermal plastids are labeled with the stroma-targeted fluorescence marker RecARed (green). These plastids also produce a lot of string-like protrusion of their envelope, called stromules. The plastids from the deeper cells are bigger and have a strong chlorophyll autofluorescence (magenta). The maximum projection image is in the galley above.
The video portrays the 3D reconstruction of the Z stack of moss gametophore "leaf" cells. We can see chloroplasts (in green) and walls between neighboring cells (in magenta). Both are visible due to their natural autofluorescence. The maximum projection image is in the galley above.